Computer scientists and biologists have identified an underlying genomic signature for 29 different COVID-19 RNA sequences using machine learning.
This new data discovery tool will help researchers to classify viruses like COVID-19 in just a few minutes””which is a process of high importance for strategic planning and mobilizing medical needs during a pandemic.
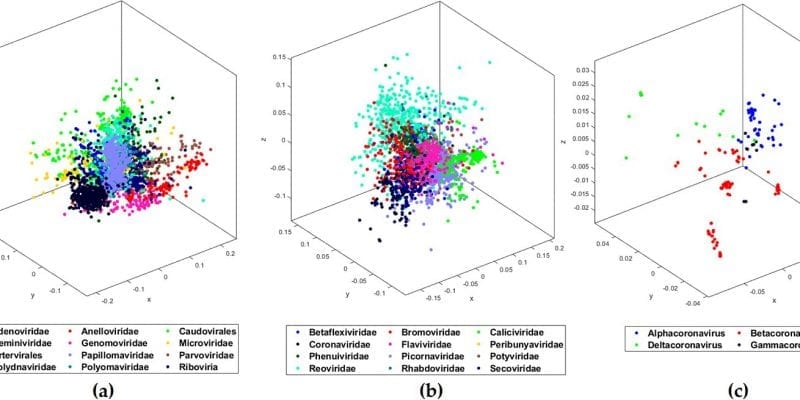
This discovery also supports the scientific hypothesis that COVID-19 (SARS-CoV-2) has its origin in bats (Sarbecovirus, a subgroup of Betacoronavirus).
The findings, Machine learning using intrinsic genomic signatures for rapid classification of novel pathogens.
The “ultra-fast, scalable, and highly accurate” classification system uses a brand-new software, graphic-based and decision-tree approach to illustrate the classification and deliver the best choice out of all possible outcomes.
Kathleen Hill, a biology professor co-led the study with Western collaborators in Statistical and Actuarial Sciences and Computer Science, along with others in the University of Waterloo’s Department of Computer Science.
This machine-learning method delivers a 100% accurate classification of the COVID-19 sequences and most importantly, discovers the most relevant relationships for over 5,000 viral genomes again within minutes.
“All we needed was the COVID-19 RNA sequence to discover its own intrinsic sequence pattern. We used that signature pattern and a logical approach to match that pattern as close as possible to other viruses and achieved a fine level of classification in minutes””not days, not hours but minutes,” Hill stated.
This classification tool has already analyzed more than 5,000 unique viral genomic sequences, including the 29 COVID-19 sequences available.
Hill believes this data discovery tool, which can classify any newly discovered virus sequence, will be an essential component for vaccine development, researchers, scientists, and health-care workers during this global pandemic and beyond.
Contact Us
Call or submit our online form to request an estimate or for general questions about our services. We look forward to serving you!